



      
|

|
Relaxation kinetics in confinement
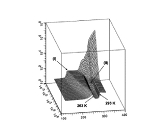
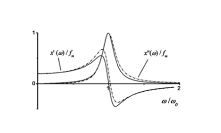
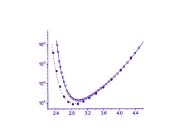
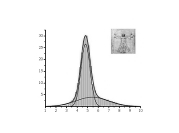
This project was originated from an unusual experimental observation which was made in the Lab of Professor Yuri Feldman. It was observed that relaxation time associated with dielectric relaxation of water molecules confined in porous glass exhibits a non-monotonic temperature dependency. At low temperatures this dependency agrees with classic Arrhenius pattern when temperature increase leads to decrease of relaxation time. However, when temperature is increased further, in this case, unlike the Arrhenius behavior, the relaxation time starts to slow down and finally returns to the initial values (see upper Figure on the right). I provided a theoretical model which can explain this type of behavior making use of some spatial confinement arguments. Later it was found that similar ideas can be useful for a number of other confined systems which we discussed in a small review on this subject. The most intriguing aspect of this story is that temperature dependency of protein folding time is also non-monotonous. I learned about this absolutely by chance when I got in my hands a paper disusing the results of some protein folding kinetics simulations. I reanalyzed those results and published a work where I treated them with my model. This exersise revealed that my model agrees with these data better than the model originally used to process them. The agreement with real experimental data was also quite satisfactory (see bottom Figure on the right). Unfortunately, this work was not really welcomed by protein folding community for the reason that it already had a pre-conception for explaining non-monotonous folding time kinetics. This explanation derives from the Random Energy Model that was borrowed from the theory of spin glasses. Still, for a number of different reasons I do believe that my approach is more adequate to this problem. Most importantly because it is not only explains temperature dependence of folding time but also can be used to resolve Levinthal's paradox and predicts algebraic scaling of folding time with protein length postulated earlier. Related articles: |